Designing Agentic AI for Pharma & Life Sciences with Graph Technology: From Complex Evidence to Explainable Insights
5 min read
In this blog, we’d like to share insights from our recent webinar on Agentic AI in Pharma and Life Sciences and how graph technology helps organizations move from fragmented data and isolated LLM experiments to scalable, explainable, and production-ready AI systems.
During the session, we discussed a recurring pattern in pharma and life sciences: complex, highly connected science is forced into rigid systems, and AI initiatives struggle because the data foundation is not designed for connectivity.
The Core Challenge: Heterogeneity, Connectivity, and Unstructured Data
Across pharma R&D, clinical, and commercial functions, we see three structural challenges:
- Heterogeneous data – gene synonyms, multiple identifiers, evolving ontologies.
- Highly connected biology – genes → RNA → proteins → pathways → diseases → drugs.
- Unstructured content at scale – publications, patents, CSRs, contracts, slide decks, interviews.
Biology is a network by definition. Yet many systems still rely on tables that require endless joins and manual stitching.
As discussed in the session, if you regularly perform 3+ joins (or combine multiple spreadsheets) to answer a question, you’re likely facing a graph problem.
A knowledge graph aligns naturally with the way scientists think: entities connected by meaningful relationships.
From Graph Thinking to Agentic AI
Graphs solve structure. Agentic AI adds intelligence.
But in regulated industries like pharma, we must avoid black-box systems. The design principle we emphasized:
Let agents think. Let tools act.
LLMs reason about intent and context. Deterministic tools, queries, extraction pipelines, and validation logic execute the action. This separation improves reliability, governance, and explainability.
As highlighted during the session:
“God does not play dice with Agents.”
— (probably) Einstein
Large language models are inherently probabilistic. In pharma and life sciences, however, we cannot rely on probability when decisions affect research programs, patient safety, or regulatory submissions. Agentic systems must therefore combine intelligent reasoning with deterministic graph-based execution to ensure repeatability and traceability.
Schema-First Knowledge Graph Construction
A practical way to ground agents is to define a clear domain schema before extraction:
# Example domain schema
node_types = ["Molecule", "Company", "Target", "Disease"]
rel_types = ["TREATS", "TARGETS", "ASSOCIATED_WITH", "IN_PIPELINE"]
patterns = [
("Molecule", "TREATS", "Disease"),
("Molecule", "TARGETS", "Target"),
("Disease", "ASSOCIATED_WITH", "Target"),
("Company", "IN_PIPELINE", "Molecule"),
]
Unstructured sources, such as publications, patents, Clinical Study Reports (CSRs), are then processed to populate this schema. Each extracted fact can link back to its original source.
This turns:
“The molecule appears relevant.”
Into:
- Normalized molecule entity
- Connected disease entity
- Defined relationship type
- Traceable source paragraph
Explainability becomes built-in, not retrofitted.
For a hands-on example of this approach using GraphRAG with Python and Neo4j, see the Github workshop repository.
GraphRAG in Action: Controlled Querying
Instead of allowing an LLM to generate arbitrary database queries, we expose curated graph tools.
For example:
MATCH (g:Gene {symbol: $geneSymbol})-[:ENCODES]->(p:Protein)
MATCH (p)-[r:INTERACTS_WITH]->(p2:Protein)
RETURN p.id AS source_protein,
p2.id AS target_protein,
r.score AS interaction_score
ORDER BY interaction_score DESC
LIMIT 25;
The agent selects the tool.
The database executes the query deterministically.
The result remains auditable.
This model supports compliance requirements and role-based access control, critical in pharma environments.
From Chat to Dashboards
Agentic AI in pharma should not live only in a chat interface.
Insights must surface inside:
- R&D dashboards
- Competitive intelligence portals
- Clinical operations cockpits
- Safety monitoring systems
With a graph foundation, dashboards become dynamic and explainable. An agent can:
- Explain why a target appears in a ranking
- Trace connections across trials and publications
- Show the exact evidence path in the graph
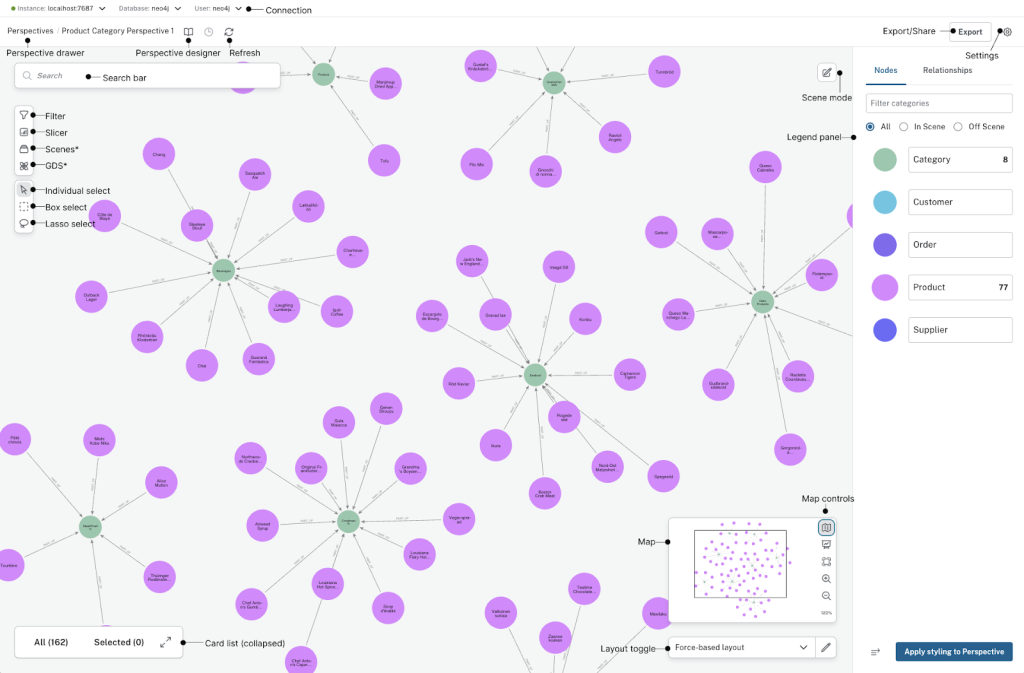
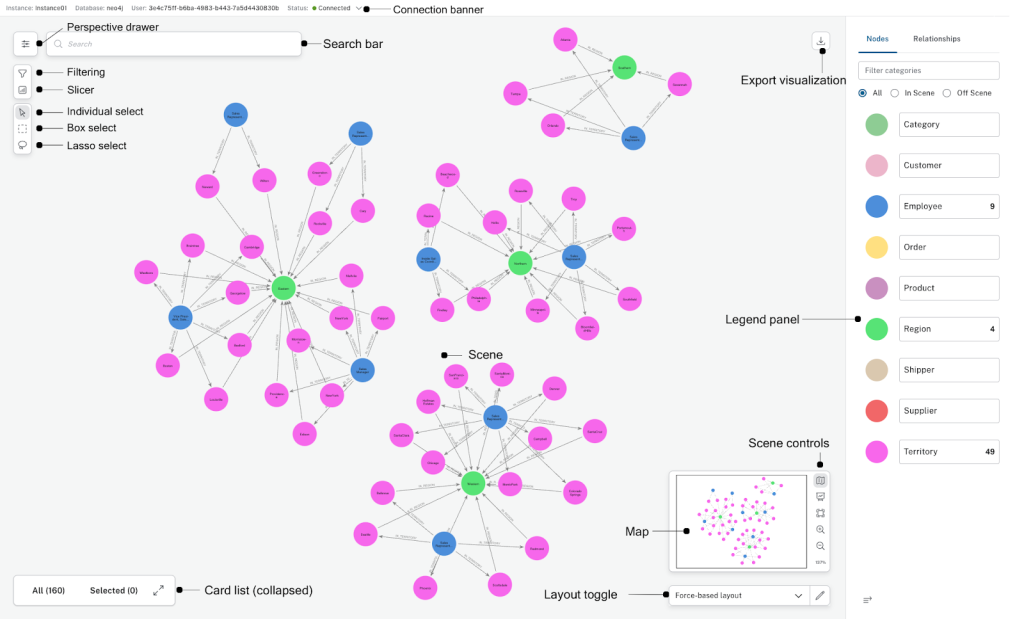
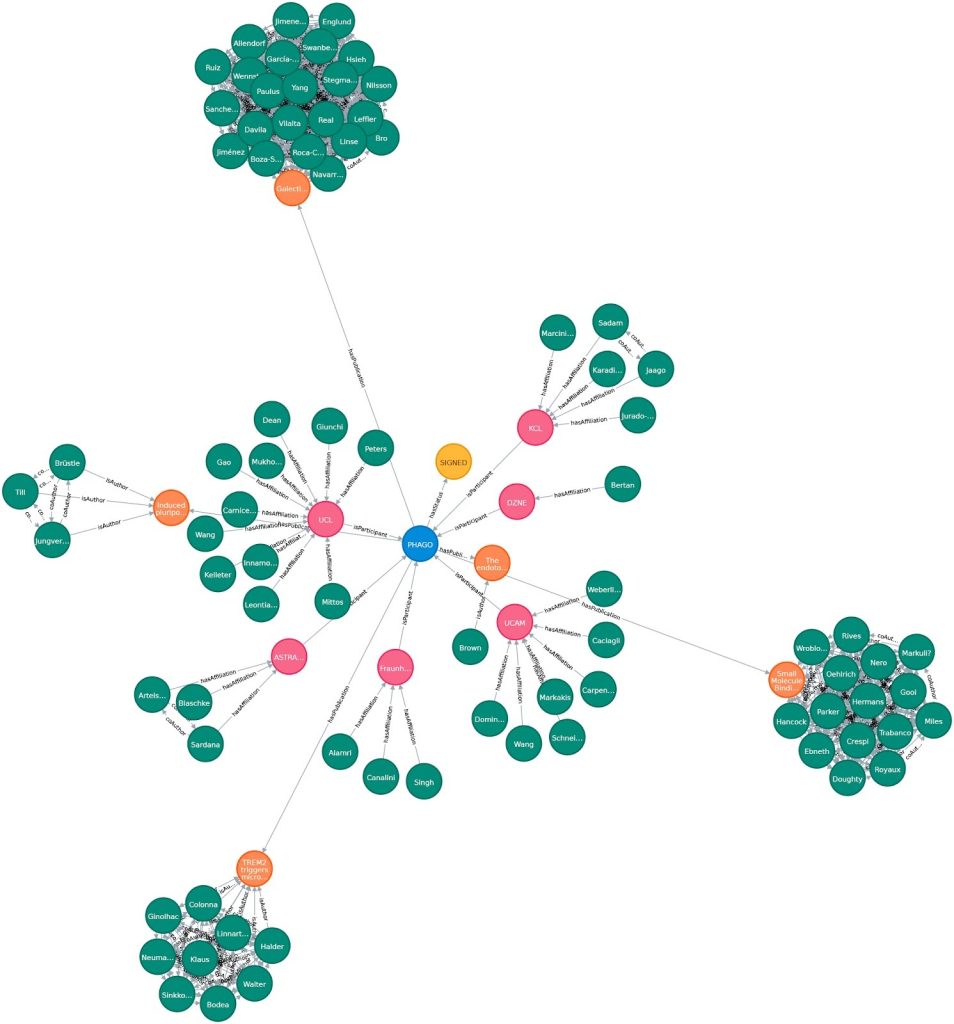
Because the visualization is backed by a knowledge graph, every metric is traceable to connected entities and relationships, not just text similarity.
Across the Drug Lifecycle
This architecture applies across the pharma value chain:
- Target identification and biomarker discovery
- Drug repurposing and competitive intelligence
- Clinical study analysis and CSR summarization
- Safety signal investigation
- Supply Chain intelligence
In each case, the graph acts as:
- A context layer for agents
- A memory layer for enterprise knowledge
- A control layer for governance and explainability
Agentic AI becomes a structured augmentation system — not a speculative assistant.
Real-World Example: Novartis and Connected Drug Discovery
This approach is already in practice. Companies like Novartis are using graph technology to accelerate early drug discovery.
In their biomedical knowledge graph initiative, Novartis integrated:
- Text-mined scientific literature
- Internal experimental datasets
- Biological background knowledge
Into a unified knowledge graph.
This enables researchers to explore gene–disease–compound relationships in a single connected system, improving biological interpretation and helping prioritize promising targets more efficiently.👉Read the full customer story
Conclusion and Next Steps
Building Agentic AI in pharma is not about adding an LLM on top of existing systems. It requires:
- A connected data foundation
- Ontology-aware extraction
- Tool-driven execution
- Explainable graph-based retrieval
Knowledge graphs provide the structure.
Agentic workflows provide scalable intelligence.
We’ll continue exploring this topic at the 3rd edition of GraphTalk Pharma & Life Sciences, a hybrid event held in Munich. In the afternoon, we’re planning a more technical session focused on building agentic systems on top of knowledge graphs and diving deeper into implementation patterns.
We look forward to continuing the conversation and building the next generation of explainable, connected AI systems together.










