BenchSci Decodes Disease Biology with Neo4j to Accelerate Drug Discovery
BenchSci transforms fragmented biomedical evidence into a graph-native model of disease biology, empowering 9 of the top 10 pharmaceutical companies to de-risk and accelerate discovery.
9 of the top 10
Pharmaceutical companies have adopted BenchSci’s technology
4,500+
Research institutions worldwide accelerate science with BenchSci’s technology
$2.5 billion
Average cost of bringing a new drug to market

Developing a single new drug takes an average of 13 years and costs over $2.5 billion. More than 90% of clinical trials fail — most often because the underlying biology was misunderstood or incompletely characterized in preclinical research.
Drug discovery is failing because disease biology remains a black box.
The diseases healthcare organizations are trying to treat today are more complex and interconnected than ever before. Diseases are multi-factorial, dynamic, and deeply interconnected. But the evidence required to understand them is scattered across millions of publications, experimental contexts, model systems, and proprietary datasets.
While scientific output continues to grow exponentially, knowledge remains fragmented. Researchers are inundated with data but lack a unified, systems-level representation of the biological evidence needed to make high-confidence decisions. BenchSci is addressing this structural failure in how science is operationalized. The company is building a computable map of disease biology: a living representation of experimentally validated biological relationships designed to accelerate and de-risk preclinical research.
ASCEND is BenchSci’s neuro-symbolic AI platform purpose-built for this challenge. At its core is the Biological Evidence Knowledge Graph (BEKG), an experimentally grounded knowledge system integrating evidence from open-access literature, closed-access publications, and proprietary pharmaceutical datasets.
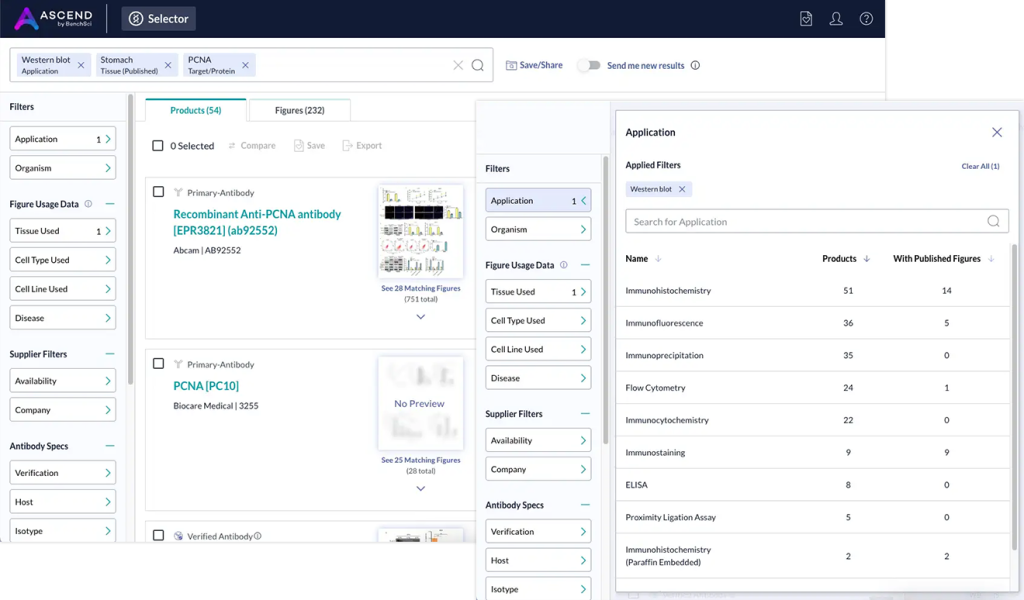
Above: ASCEND, BenchSci’s neuro-symbolic AI platform
Unlike pure generative AI systems, ASCEND does not treat biology as free text. In high-stakes scientific domains, hallucinated outputs are liabilities. ASCEND combines large language models (LLMs) with a graph-native, ontologically structured representation of biological entities and their experimentally validated relationships. Every assertion is anchored to source evidence, and every inference is traceable.
BenchSci enables scientists to generate hypotheses with greater speed, credibility, and precision by transforming fragmented scientific context into a connected, evidence-grounded intelligence layer.
The Challenge: When Biology Outgrows Rows and Columns
BenchSci confronted a fundamental modeling problem: biology does not conform to rows and columns.
Disease biology is inherently networked. A single protein may participate in multiple pathways, influence diverse phenotypes, and exhibit context-dependent behavior across experimental systems.
“You’re dealing with billions of entities and a combinatorial explosion of possible interactions,” says Dr. Tony Solon, VP of Engineering, Data & ML at BenchSci. “But more importantly, you’re dealing with relationships that are conditional, contextual, and experimentally defined. Biology only makes sense when those connections are preserved.”
Early architectures relied on relational databases and denormalized tables. While sufficient for structured metadata, they proved brittle when modeling deeply interconnected evidence extracted from millions of publications. Multi-hop biological reasoning required increasingly complex JOIN operations, and granular subgraphs derived from literature became fragmented islands of information.
Critically, there was no native linkage between a structured assertion and its originating experiment. Claims extracted from PDFs became detached from the evidentiary context required to interpret them.
“We realized we were facing both a performance and a representational issue,” Dr. Solon explains. “If relationships are first-class citizens in biology, they must be first-class citizens in the data model. Without that, you lose traceability, explainability, and ultimately scientific credibility.”
BenchSci required an architecture built around relationships to enable scientists to ask complex, multi-hop questions, such as identifying protein targets associated with a disease while minimizing downstream toxicity risk.
The Solution: A Neuro-symbolic Brain Built on Neo4j
BenchSci built the Biological Evidence Knowledge Graph (BEKG) on Neo4j to operationalize a computable model of disease biology. This graph-native foundation is designed to preserve biological relationships at scale.
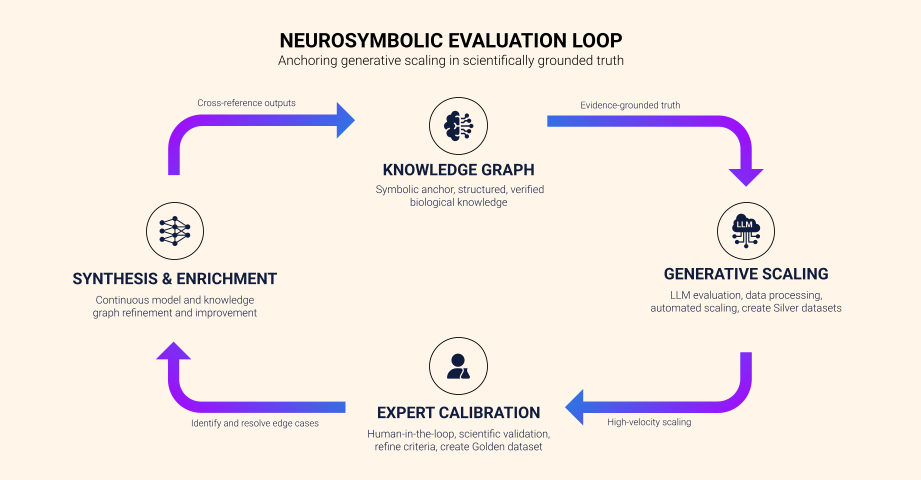
Above: BenchSci’s neuro-symbolic evaluation loop
The BEKG contains hundreds of millions of biological entities and relationships derived from evidence: tens of millions of scientific publications and proprietary datasets. The architecture was designed to ensure that biological reasoning remains structurally intact.
BenchSci’s approach is explicitly neuro-symbolic. It combines large language models with a formally structured knowledge graph, addressing the limitations of both pure generative AI — which can hallucinate — and purely symbolic systems, which can become brittle and incomplete.
1. LENS: Reading Like a Scientist
During knowledge extraction, BenchSci’s Layered Extraction Network Semantics (LENS) engine uses Google Cloud’s Gemini models and proprietary vision systems to interpret scientific papers with contextual awareness. Unlike conventional text mining, LENS distinguishes whether a biological term (e.g., p53) is functioning as a target, reagent, biomarker, or outcome variable within a specific experimental setup. It also validates claims against associated experimental imagery, such as Western blots and microscopy data. This ensures that extracted relationships are not merely textual co-occurrences, but experimentally grounded assertions.
2. Ontological Governance: T-Box and A-Box Separation
The BEKG is governed by a rigorous ontological framework implemented in Neo4j.
BenchSci maintains a formal T-Box (Terminological Box) that defines the schema and permissible relationships between biological entities. Extracted assertions populate the A-Box (Assertional Box), which contains the billions of evidence-backed data points derived from literature and proprietary sources.
“Integrating heterogeneous biological data into a unified system is an ontological challenge,” says Dr. Solon. “Every extracted assertion must conform to a formally defined schema. That constraint is what allows us to reason over the graph with confidence. Without ontological governance, explainability and traceability break down.”
By separating schema from assertion, BenchSci ensures structural consistency while allowing the knowledge graph to scale dynamically as new evidence emerges.
3. Grounding Generative AI in Structured Evidence
Neo4j functions as the evidentiary backbone for ASCEND’s generative capabilities.
When a scientist submits a natural language query, the system first retrieves a contextually relevant subgraph from the BEKG. Only then does the generative model synthesize a response. This retrieval-augmented pattern ensures outputs are constrained by experimentally validated relationships rather than unconstrained language generation.
“It’s not sufficient to have a rich dataset,” Dr. Solon explains. “Scientific inference requires a sound semantic model. The graph provides the structure against which reasoning occurs. That’s what enables full traceability from generated insight back to the source experiment.”
Why Neo4j?
BenchSci selected Neo4j because modeling disease biology requires a graph-native architecture.
Biological reasoning is inherently multi-hop and path-dependent. Answering a single scientific question may require traversing from a target to interacting proteins, to pathways, to phenotypes, to disease contexts—all while preserving experimental provenance. In relational systems, these queries demand increasingly complex JOIN operations that degrade in performance and maintainability as the network grows.
Neo4j’s index-free adjacency enables deep traversals across billions of relationships with predictable performance. This allows BenchSci to retrieve biologically meaningful subgraphs in milliseconds, even as the Biological Evidence Knowledge Graph scales to hundreds of millions of entities.
“We have an ever-expanding number of entry points into the graph,” says Dr. Solon. “Some originate from scientists exploring hypotheses. Others come from automated agents orchestrating inference workflows. Neo4j allows us to resolve those queries against a deeply connected biological network without compromising performance, traceability, or structural integrity.”
Neo4j supports BenchSci’s ontological governance and graph analytics workflows, including graph embeddings and semantic reasoning. As the BEKG evolves, Neo4j provides the flexibility required to extend the schema, incorporate new evidence types, and compute higher-order relationships across the biological network.
“The partnership has been instrumental,” Dr. Solon adds. “Neo4j gives us the graph foundation necessary to operationalize neuro-symbolic AI at enterprise scale.”
Built for Scale on Google Cloud
BenchSci operates as a cloud-native organization on Google Cloud Platform (GCP), leveraging its infrastructure to power large-scale multimodal model training and deployment.
Vertex AI supports the deployment and orchestration of multimodal foundation models that interpret scientific text, tables, and imagery within a unified extraction pipeline. This provides the computational backbone required to process millions of scientific documents with contextual precision.
But compute alone does not solve biological reasoning.
Unstructured PDFs are ingested and processed within GCP, where LENS extracts experimentally grounded assertions. These structured outputs are then materialized into Neo4j, which serves as the persistent knowledge layer for biological inference.
BigQuery supports large-scale analytical workflows across the corpus, while Neo4j enables real-time, multi-hop traversal and reasoning across the Biological Evidence Knowledge Graph. This architectural separation allows BenchSci to expand continuously while preserving structural integrity, provenance, and enterprise-grade performance.
As biomedical research output accelerates, this foundation ensures that growth in data volume enhances, rather than degrades, the fidelity of biological insight.
Impact: From Search to Inference
The transition from keyword-based search to graph-native inference has produced measurable impact across BenchSci’s enterprise customers.
ASCEND is now deployed by 9 of the top 10 global pharmaceutical companies, including Sanofi and Merck, as well as more than 4,500 research institutions worldwide. For these organizations, the shift is not incremental—it fundamentally changes how biological decisions are made.
De-risking Portfolios: By exposing the full relational context of disease pathways, scientists can identify mechanistic gaps, conflicting evidence, and downstream liabilities earlier in the discovery process. This reduces investment in targets that lack robust biological support and lowers the probability of late-stage failure.
Scientific Productivity: Researchers spend significantly less time manually reviewing literature and more time designing experiments. Instead of aggregating evidence, they can interrogate a structured, provenance-aware knowledge graph.
Financial Efficiency: By surfacing experimentally validated performance data on reagents — including antibodies with documented reliability issues — BenchSci helps mitigate the financial and scientific cost of irreproducible research.
BenchSci is moving beyond retrieval into predictive biological reasoning.
Through its collaboration with Mila (Quebec AI Institute), the company is advancing graph machine learning models that operate directly on the Biological Evidence Knowledge Graph. These models analyze network topology, relational patterns, and latent structure to infer plausible biological relationships that have not yet been explicitly reported in the literature.
Rather than simply retrieving known associations, the system can predict novel gene–disease links or pathway interactions based on graph structure, generating high-confidence hypotheses for experimental validation.
This evolution marks a shift from AI-assisted search to AI-guided discovery. BenchSci is progressing toward a future in which hypothesis generation, evaluation, and refinement occur within a continuous, evidence-constrained feedback loop.
The Future: The Lab-in-the-Loop
BenchSci has partnered with Thermo Fisher Scientific to integrate its biological intelligence and AI capabilities with Thermo Fisher’s automation and instrument ecosystem to create a unified Lab-in-the-Loop system where computational reasoning and physical execution operate as one continuous, learning platform.
This model will eliminate the traditional divide between the dry lab and the wet lab, ensuring that digital insight flows directly into automated execution, and experimental outcomes immediately strengthen digital intelligence.
“We’re building toward a system where experimental design is informed by a continuously evolving, evidence-grounded model of biology,” says Dr. Solon. “The graph provides the structural memory of the system. As new data enters, it reshapes the knowledge landscape and informs what should be tested next.”
This Lab-in-the-Loop paradigm elevates the BEKG from organizing biological knowledge to actively shaping it by filling gaps in the existing knowledge and solidifying the understanding of biological “truth” through iterative experimental feedback.
This autonomous flywheel between structured biological knowledge and physical experimentation will enable a future where hypothesis generation, validation, and refinement occur at scale, while remaining grounded in experimental evidence.
“We provide scientists with a high-fidelity representation of disease biology,” Dr. Solon concludes. “In drug discovery, decisions routinely involve millions of dollars and the potential to impact patient lives. Speed is critical. But credibility and reliability in the underlying intelligence system are non-negotiable.”
