Network science: The hidden field behind machine learning, economics and genetics that you’ve (probably) never heard of – an interview with Dr. Aaron Clauset [Part 1]

Graph Analytics & AI Program Director
10 min read

I recently had the opportunity to combine work and pleasure and meet with Dr. Aaron Clauset, an expert on network science, data science and complex systems. In 2016, Clauset won the Erdos-Renyi Prize in Network Science but you might be more familiar with his earlier research into power-laws, link prediction and modularity.
Dr. Clauset directs the research group that developed the ICON dataset reference (if you’re looking for network data to test, bookmark this now) and has recently published research that sheds light on possible misconceptions about network structures. When a last-minute business trip to Denver came up, I made the trip up to Boulder where Clauset is an assistant professor of computer science at University of Colorado Boulder.
Dr. Aaron Clauset is an Assistant Professor of Computer Science at the University of Colorado Boulder and in the BioFrontiers Institute. He’s also part of the external faculty at the Santa Fe Institute (for complexity studies).
Between lunch and Clauset’s next class, we chatted about his group’s recent research and the general direction of network science, and I left with a superposition of both disillusionment and excitement. The Clauset Lab has been working to expand the diversity and rigor of studying complex systems and in doing so, they may be dismantling some cherished beliefs that date back to the 90s. (I should have known it wouldn’t be simple; we’re talking about complex systems after all.)
This matters to the Neo4j graph community because anyone analyzing networks – especially if they are looking for global attributes – needs to understand the underlying dynamics and structure. Below is a summary of our discussion.
What kind of work is your team focused on?
Clauset: My research group at CU Boulder currently includes five Ph.D. students, along with a few masters and several undergraduates. Our research focuses on both developing novel computational methods for understanding messy, complicated datasets, and on applying these methods to solve real scientific problems, in biological and social settings mainly.
In the group, everyone is involved in research in some way. For example, the ICON website (index of complex networks) was built by a pair of undergrads to teach themselves networks concepts and explore tools.
Networks are one of our key areas of work. Networks are really just a representation, a tool to understand complex systems. We represent how a social system works by thinking about interactions between pairs of people. By analyzing the structure of this representation, we can then answer questions about how the system works or how individuals behave within it. In this sense, network science is a set of technical tools that can be applied to nearly any domain.
Networks also act as a bridge for understanding how microscopic interactions and dynamics can lead to global or macroscopic regularities. They can bridge between the micro and the macro because they represent exactly which things are interacting with each other. It used to be common to assume that everything interacts with everything, and we know that’s not true; in genetics, not all pairs of people and not all pairs of genes interact with each other.
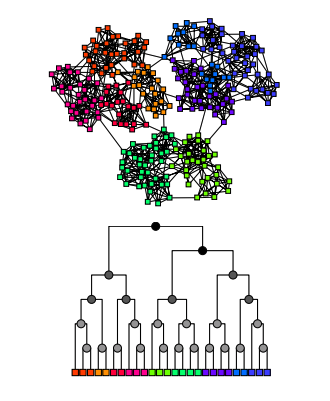
Taken from, “Hierarchical structure and the prediction of missing links in networks“
An extremely important effort in network science is figuring out how the structure of a network shapes the dynamics of the whole system. We’ve learned in the last 15 years that for many complex systems, the network is incredibly important in shaping both what happens to individuals within the network and how the whole system evolves.
My group’s work focuses on characterizing the structure of these networks so that we can better understand how structure ultimately shapes function.
Are there commonalities across different types of networks?
Clauset: In the late 1990s and early 2000s, a lot of energy in driving network science came from physicists, who brought new mathematical tools, models and lots of new data. One idea they popularized was the hypothesis that “universal” patterns occurred in networks of all different kinds – social, biological, technological, information and even economic networks – and these were driven by a small number of fundamental processes.
This kind of idea was pretty normal in one part of physics. For instance, there’s a universal mathematical model of how a magnet works that makes remarkably accurate predictions about real magnets of all different kinds.
The dream for networks was to show that the same could be done for them: that all different kinds of networks could be explained by a small set of basic mathematical principles or processes, or that they fell into a small number of general structural categories. It’s a pretty powerful idea, and it inspired both a lot of really good, cross-disciplinary work, as well as a number of highly provocative claims.
![]() The validity of some of the boldest claims has been difficult to evaluate empirically because it required using a large and diverse set of real-world networks to test empirical “universality” of the pattern. Assembling such a dataset is part of what led us to put together the Index of Complex Networks, what we call the ICON index.
The validity of some of the boldest claims has been difficult to evaluate empirically because it required using a large and diverse set of real-world networks to test empirical “universality” of the pattern. Assembling such a dataset is part of what led us to put together the Index of Complex Networks, what we call the ICON index.
Although we’re still expanding it, my group has already begun revisiting many of the early claims about universal patterns in networks, including the idea that “all networks are scale-free,” or that only social networks have a high triangle density, or that networks cluster into “superfamilies” based on the pattern of their local structure. Surprisingly, many claims about the structure of networks have been repeated again and again in the literature, but they haven’t been carefully scrutinized with empirical data.
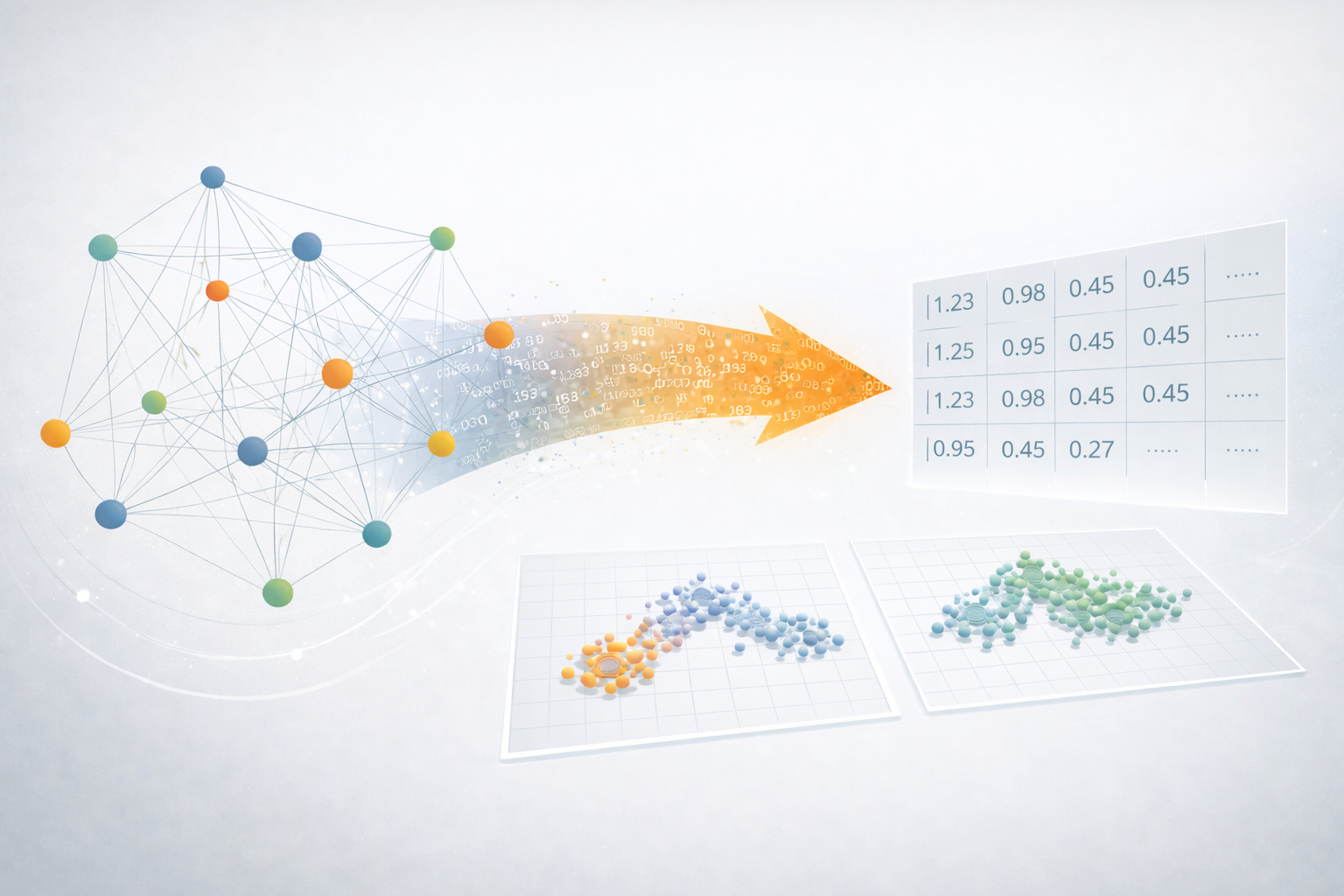
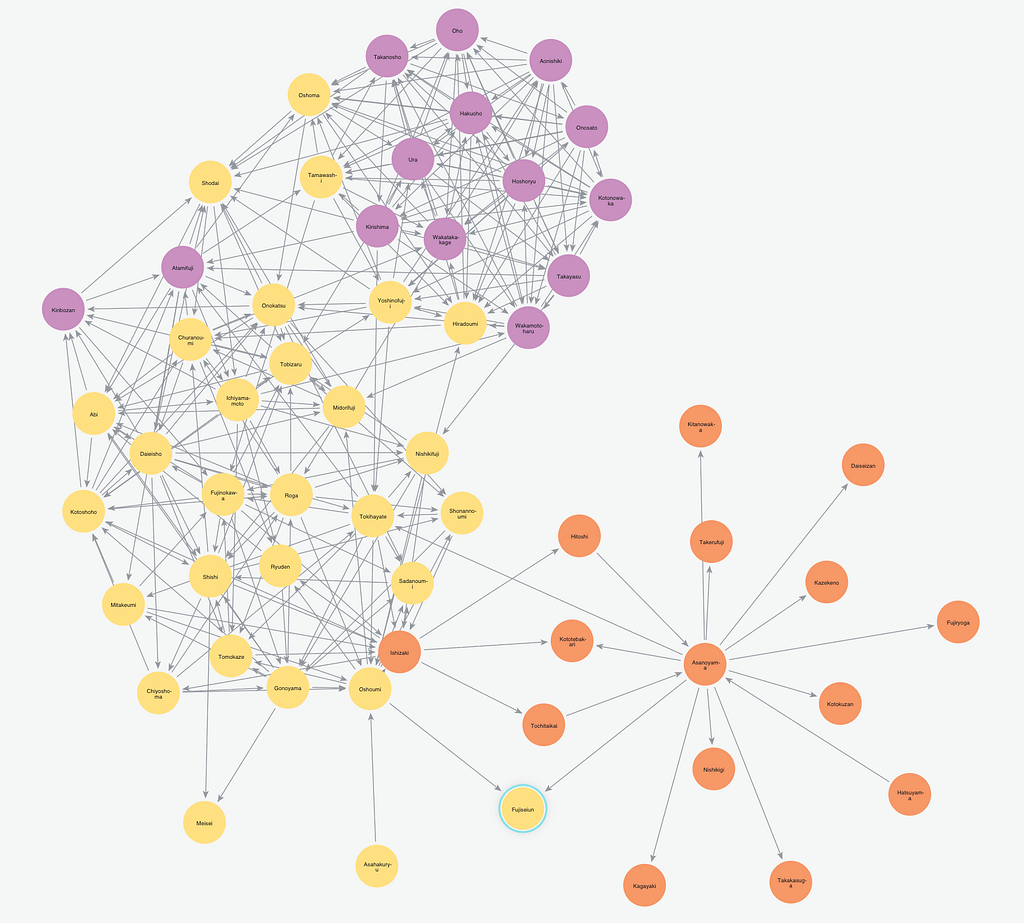
It turns out that many of these universal patterns fall apart when you can look across a huge variety of networks. Kansuke Ikehara’s recent paper [Characterizing the structural diversity of complex networks across domains] asks a simple question: If I label a large number of networks with where they came from (for example a transportation/road network, an online/social network, or a metabolic/biological network) can you use machine learning to discover what features distinguish these classes of networks?
 If there are a few “families” of network structures, then no algorithm should be able to learn to distinguish different networks within a family. Instead, what we found was that pretty much every class of network was easily distinguishable from every other class.
If there are a few “families” of network structures, then no algorithm should be able to learn to distinguish different networks within a family. Instead, what we found was that pretty much every class of network was easily distinguishable from every other class.
Social networks cluster together in one part of the feature space, biological networks are generally well separated from those, etc., and this is true for every class of network we looked at. The clear take-home message is that there’s a lot more diversity in network structures than we thought 20 years ago, and therefore a lot more work to be done to understand where this diversity comes from.
Ikehara’s research revealed the hidden structural diversity of networks and suggests that there may be fewer universal patterns than once thought. At the same time, some clusters of networks are closer to each other in terms of their structure.
For instance, we found that water distribution networks exhibit similar structural signatures to fungal mycelia networks, which suggests that they may be shaped by similar underlying processes or optimization problems. In this way, machine learning can help us identify such structural commonalities, and therefore help us figure out, in a data-driven way, where we might most likely to find a common mechanistic explanation.
How is network science evolving?
Clauset: In many ways network science today is diversifying and expanding. This expansion is enabling a great deal of specialization, but there’s a trade-off. People can now take network methods and apply them in really specific questions about really specific systems.
This is enormously productive and an exciting achievement for network science. But, the growth of disciplinary work around networks also means there’s relatively less work that crosses disciplinary boundaries. Without shared spaces where people from different domains get together to talk about their breakthroughs, people working on one type of problem are less likely to get exposed to potentially remarkable ideas in a different area.
Sure, a lot of ideas about economics won’t apply to biological networks, but some will, and if the economists and biologists never talk to each other, we’ll never know. If there’s no common ground, there will be a lot of reinvention and delays, even years for methods in one domain to cross over to another.
This is why I think it’s so important to study and get together to discuss networks in general. This kind of cross-disciplinary fervor is another thing physicists and computer scientists helped get going about 20 years ago; it was mainly physicists and computer scientists broadcasting “we can do sociology, and politics, and ecology too.”
That attitude certainly annoyed some people, especially the sociologists who’d been doing networks for 80 years already, but it also generated enormous and broad interest in networks across essentially all the sciences. Now, the different disciplinary areas of network science are growing so quickly that in some ways the middle – the crossroads where ideas can jump between fields – is effectively shrinking.
How can network science encourage more cross-domain collaboration?
 Clauset: Having an actual event that serves as a crossroads between domains where people can present and interact is essential. In many ways the International Conference on Network Science is trying to do that, but it struggles to pull researchers out of their domains and into the middle, since different disciplines have different overarching questions. I think so long as some domain experts from different fields come to the crossroads to talk and interact, good ideas will eventually spread.
Clauset: Having an actual event that serves as a crossroads between domains where people can present and interact is essential. In many ways the International Conference on Network Science is trying to do that, but it struggles to pull researchers out of their domains and into the middle, since different disciplines have different overarching questions. I think so long as some domain experts from different fields come to the crossroads to talk and interact, good ideas will eventually spread.
Continuing this interdisciplinary effort will be a key part of continuing the advancement of network science. But, not all efforts need to be interdisciplinary. In fact, disciplines are essential to help focus our collective attention.
I’m not sure what the right balance is between disciplinary and interdisciplinary work, but for me, interdisciplinary ideas are the most exciting. If work on these isn’t funded and supported at decent levels, we surely won’t address many of the most important ideas in society because they’re the ones that span different disciplines.
For instance, cybersecurity is not just a technical issue because humans have a terrible track record of writing bug-free software. Real security requires legal components, social components, ethical components, economic components, and probably more to develop a lasting solution.
In fact, if you choose any problem that impacts a decent portion of the population, then it’s surely an interdisciplinary problem that will require an interdisciplinary approach to understand and solve.
Conclusion
As you can see, we had a great discussion on how some of the preconceived ideas around networks are changing. Next week – in the second part of this series – I’ll summarize our deeper dive into some of the advancements and emerging themes in network science.