Closeness Centrality
Glossary
- Directed
-
Directed trait. The algorithm is well-defined on a directed graph.
- Directed
-
Directed trait. The algorithm ignores the direction of the graph.
- Directed
-
Directed trait. The algorithm does not run on a directed graph.
- Undirected
-
Undirected trait. The algorithm is well-defined on an undirected graph.
- Undirected
-
Undirected trait. The algorithm ignores the undirectedness of the graph.
- Heterogeneous nodes
-
Heterogeneous nodes fully supported. The algorithm has the ability to distinguish between nodes of different types.
- Heterogeneous nodes
-
Heterogeneous nodes allowed. The algorithm treats all selected nodes similarly regardless of their label.
- Heterogeneous relationships
-
Heterogeneous relationships fully supported. The algorithm has the ability to distinguish between relationships of different types.
- Heterogeneous relationships
-
Heterogeneous relationships allowed. The algorithm treats all selected relationships similarly regardless of their type.
- Weighted relationships
-
Weighted trait. The algorithm supports a relationship property to be used as weight, specified via the relationshipWeightProperty configuration parameter.
- Weighted relationships
-
Weighted trait. The algorithm treats each relationship as equally important, discarding the value of any relationship weight.
- Node properties
-
Node properties trait. The algorithm makes use of node properties.
Introduction
Closeness centrality is a way of detecting nodes that are able to spread information very efficiently through a graph.
The closeness centrality of a node measures its average farness (inverse distance) to all other nodes. Nodes with a high closeness score have the shortest distances to all other nodes.
For each node u, the Closeness Centrality algorithm calculates the sum of its distances to all other nodes, based on calculating the shortest paths between all pairs of nodes. The resulting sum is then inverted to determine the closeness centrality score for that node.
The raw closeness centrality of a node u is calculated using the following formula:
raw closeness centrality(u) = 1 / sum(distance from u to all other nodes)
It is more common to normalize this score so that it represents the average length of the shortest paths rather than their sum. This adjustment allow comparisons of the closeness centrality of nodes of graphs of different sizes
The formula for normalized closeness centrality of node u is as follows:
normalized closeness centrality(u) = (number of nodes - 1) / sum(distance from u to all other nodes)
Wasserman and Faust have proposed an improved formula for dealing with unconnected graphs. Assuming that n is the number of nodes reachable from u (counting also itself), their corrected formula for a given node u is given as follows:
Wasserman-Faust normalized closeness centrality(u) = (n-1)^2/ number of nodes - 1) * sum(distance from u to all other nodes
Note that in the case of a directed graph, closeness centrality is defined alternatively. That is, rather than considering distances from u to every other node, we instead sum and average the distance from every other node to u.
Use-cases - when to use the Closeness Centrality algorithm
-
Closeness centrality is used to research organizational networks, where individuals with high closeness centrality are in a favourable position to control and acquire vital information and resources within the organization. One such study is "Mapping Networks of Terrorist Cells" by Valdis E. Krebs.
-
Closeness centrality can be interpreted as an estimated time of arrival of information flowing through telecommunications or package delivery networks where information flows through shortest paths to a predefined target. It can also be used in networks where information spreads through all shortest paths simultaneously, such as infection spreading through a social network. Find more details in "Centrality and network flow" by Stephen P. Borgatti.
-
Closeness centrality has been used to estimate the importance of words in a document, based on a graph-based keyphrase extraction process. This process is described by Florian Boudin in "A Comparison of Centrality Measures for Graph-Based Keyphrase Extraction".
Constraints - when not to use the Closeness Centrality algorithm
-
Academically, closeness centrality works best on connected graphs. If we use the original formula on an unconnected graph, we can end up with an infinite distance between two nodes in separate connected components. This means that we’ll end up with an infinite closeness centrality score when we sum up all the distances from that node.
In practice, a variation on the original formula is used so that we don’t run into these issues.
Syntax
This section covers the syntax used to execute the Closeness Centrality algorithm in each of its execution modes. We are describing the named graph variant of the syntax. To learn more about general syntax variants, see Syntax overview.
CALL gds.closeness.stream(
graphName: String,
configuration: Map
)
YIELD
nodeId: Integer,
score: Float| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
graphName |
String |
|
no |
The name of a graph stored in the catalog. |
configuration |
Map |
|
yes |
Configuration for algorithm-specifics and/or graph filtering. |
| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
List of String |
|
yes |
Filter the named graph using the given node labels. Nodes with any of the given labels will be included. |
|
List of String |
|
yes |
Filter the named graph using the given relationship types. Relationships with any of the given types will be included. |
|
Integer |
|
yes |
The number of concurrent threads used for running the algorithm. |
|
String |
|
yes |
An ID that can be provided to more easily track the algorithm’s progress. |
|
Boolean |
|
yes |
If disabled the progress percentage will not be logged. |
|
useWassermanFaust |
Boolean |
|
yes |
Use the improved Wasserman-Faust formula for closeness computation. |
1. In a GDS Session, the default is the number of available processors. |
||||
| Name | Type | Description |
|---|---|---|
nodeId |
Integer |
Node ID. |
score |
Float |
Closeness centrality score. |
CALL gds.closeness.stats(
graphName: String,
configuration: Map
)
YIELD
centralityDistribution: Map,
computeMillis: Integer,
postProcessingMillis: Integer,
preProcessingMillis: Integer,
configuration: Map| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
graphName |
String |
|
no |
The name of a graph stored in the catalog. |
configuration |
Map |
|
yes |
Configuration for algorithm-specifics and/or graph filtering. |
| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
List of String |
|
yes |
Filter the named graph using the given node labels. Nodes with any of the given labels will be included. |
|
List of String |
|
yes |
Filter the named graph using the given relationship types. Relationships with any of the given types will be included. |
|
Integer |
|
yes |
The number of concurrent threads used for running the algorithm. |
|
String |
|
yes |
An ID that can be provided to more easily track the algorithm’s progress. |
|
Boolean |
|
yes |
If disabled the progress percentage will not be logged. |
|
useWassermanFaust |
Boolean |
|
yes |
Use the improved Wasserman-Faust formula for closeness computation. |
2. In a GDS Session, the default is the number of available processors. |
||||
| Name | Type | Description |
|---|---|---|
centralityDistribution |
Map |
Map containing min, max, mean as well as p50, p75, p90, p95, p99 and p999 percentile values of centrality values. |
preProcessingMillis |
Integer |
Milliseconds for preprocessing the graph. |
computeMillis |
Integer |
Milliseconds for running the algorithm. |
postProcessingMillis |
Integer |
Milliseconds for computing the statistics. |
configuration |
Map |
Configuration used for running the algorithm. |
CALL gds.closeness.mutate(
graphName: String,
configuration: Map
)
YIELD
nodePropertiesWritten: Integer,
preProcessingMillis: Integer,
computeMillis: Integer,
postProcessingMillis: Integer,
mutateMillis: Integer,
centralityDistribution: Map,
configuration: Map| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
graphName |
String |
|
no |
The name of a graph stored in the catalog. |
configuration |
Map |
|
yes |
Configuration for algorithm-specifics and/or graph filtering. |
| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
mutateProperty |
String |
|
no |
The node property in the GDS graph to which the centrality is written. |
List of String |
|
yes |
Filter the named graph using the given node labels. |
|
List of String |
|
yes |
Filter the named graph using the given relationship types. |
|
Integer |
|
yes |
The number of concurrent threads used for running the algorithm. |
|
Boolean |
|
yes |
If disabled the progress percentage will not be logged. |
|
String |
|
yes |
An ID that can be provided to more easily track the algorithm’s progress. |
|
useWassermanFaust |
Boolean |
|
yes |
Use the improved Wasserman-Faust formula for closeness computation. |
| Name | Type | Description |
|---|---|---|
nodePropertiesWritten |
Integer |
Number of properties added to the in-memory graph. |
preProcessingMillis |
Integer |
Milliseconds for preprocessing the graph. |
computeMillis |
Integer |
Milliseconds for running the algorithm. |
postProcessingMillis |
Integer |
Milliseconds for computing the statistics. |
mutateMillis |
Integer |
Milliseconds for mutating the GDS graph. |
centralityDistribution |
Map |
Map containing min, max, mean as well as p50, p75, p90, p95, p99 and p999 percentile values of centrality values. |
configuration |
Map |
Configuration used for running the algorithm. |
CALL gds.closeness.write(
graphName: String,
configuration: Map
)
YIELD
nodePropertiesWritten: Integer,
preProcessingMillis: Integer,
computeMillis: Integer,
postProcessingMillis: Integer,
writeMillis: Integer,
centralityDistribution: Map,
configuration: Map| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
graphName |
String |
|
no |
The name of a graph stored in the catalog. |
configuration |
Map |
|
yes |
Configuration for algorithm-specifics and/or graph filtering. |
| Name | Type | Default | Optional | Description |
|---|---|---|---|---|
List of String |
|
yes |
Filter the named graph using the given node labels. Nodes with any of the given labels will be included. |
|
List of String |
|
yes |
Filter the named graph using the given relationship types. Relationships with any of the given types will be included. |
|
Integer |
|
yes |
The number of concurrent threads used for running the algorithm. |
|
String |
|
yes |
An ID that can be provided to more easily track the algorithm’s progress. |
|
Boolean |
|
yes |
If disabled the progress percentage will not be logged. |
|
Integer |
|
yes |
The number of concurrent threads used for writing the result to Neo4j. |
|
String |
|
no |
The node property in the Neo4j database to which the centrality is written. |
|
useWassermanFaust |
Boolean |
|
yes |
Use the improved Wasserman-Faust formula for closeness computation. |
3. In a GDS Session, the default is the number of available processors. |
||||
| Name | Type | Description |
|---|---|---|
nodePropertiesWritten |
Integer |
Number of properties written to Neo4j. |
preProcessingMillis |
Integer |
Milliseconds for preprocessing the graph. |
computeMillis |
Integer |
Milliseconds for running the algorithm. |
postProcessingMillis |
Integer |
Milliseconds for computing the statistics. |
writeMillis |
Integer |
Milliseconds for mutating the GDS graph. |
centralityDistribution |
Map |
Map containing min, max, mean as well as p50, p75, p90, p95, p99 and p999 percentile values of centrality values. |
configuration |
Map |
Configuration used for running the algorithm. |
Examples
|
All the examples below should be run in an empty database. The examples use Cypher projections as the norm. Native projections will be deprecated in a future release. |
In this section we will show examples of running the Closeness Centrality algorithm on a concrete graph. The intention is to illustrate what the results look like and to provide a guide in how to make use of the algorithm in a real setting. We will do this on a small sample graph of a handful nodes connected in a particular pattern. The example graph looks like this:
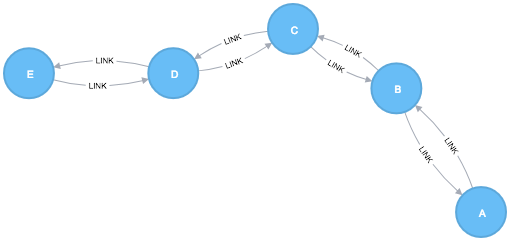
CREATE (a:Node {id:"A"}),
(b:Node {id:"B"}),
(c:Node {id:"C"}),
(d:Node {id:"D"}),
(e:Node {id:"E"}),
(a)-[:LINK]->(b),
(b)-[:LINK]->(a),
(b)-[:LINK]->(c),
(c)-[:LINK]->(b),
(c)-[:LINK]->(d),
(d)-[:LINK]->(c),
(d)-[:LINK]->(e),
(e)-[:LINK]->(d);With the graph in Neo4j we can now project it into the graph catalog to prepare it for algorithm execution.
We do this using a Cypher projection targeting the Node nodes and the LINK relationships.
MATCH (source:Node)-[r:LINK]->(target:Node)
RETURN gds.graph.project(
'myGraph',
source,
target
)In the following examples we will demonstrate using the Closeness Centrality algorithm on this graph.
Memory Estimation
First off, we will estimate the cost of running the algorithm using the estimate procedure.
This can be done with any execution mode.
We will use the stream mode in this example.
Estimating the algorithm is useful to understand the memory impact that running the algorithm on your graph will have.
When you later actually run the algorithm in one of the execution modes the system will perform an estimation.
If the estimation shows that there is a very high probability of the execution going over its memory limitations, the execution is prohibited.
To read more about this, see Automatic estimation and execution blocking.
For more details on estimate in general, see Memory Estimation.
CALL gds.closeness.stream.estimate('myGraph',{})
YIELD nodeCount, relationshipCount, bytesMin, bytesMax, requiredMemory| nodeCount | relationshipCount | bytesMin | bytesMax | requiredMemory |
|---|---|---|---|---|
5 |
8 |
1504 |
1504 |
"1504 Bytes" |
Stream
In the stream execution mode, the algorithm returns the centrality for each node.
This allows us to inspect the results directly or post-process them in Cypher without any side effects.
For example, we can order the results to find the nodes with the highest closeness centrality.
For more details on the stream mode in general, see Stream.
stream mode:CALL gds.closeness.stream('myGraph')
YIELD nodeId, score
RETURN gds.util.asNode(nodeId).id AS id, score
ORDER BY score DESC| id | score |
|---|---|
"C" |
0.6666666666666666 |
"B" |
0.5714285714285714 |
"D" |
0.5714285714285714 |
"A" |
0.4 |
"E" |
0.4 |
C is the best connected node in this graph, although B and D aren’t far behind. A and E don’t have close ties to many other nodes, so their scores are lower. Any node that has a direct connection to all other nodes would score 1.
Stats
In the stats execution mode, the algorithm returns a single row containing a summary of the algorithm result.
This execution mode does not have any side effects.
It can be useful for evaluating algorithm performance by inspecting the computeMillis return item.
In the examples below we will omit returning the timings.
The full signature of the procedure can be found in the syntax section.
For more details on the stats mode in general, see Stats.
stats mode:CALL gds.closeness.stats('myGraph')
YIELD centralityDistribution
RETURN centralityDistribution.min AS minimumScore, centralityDistribution.mean AS meanScore| minimumScore | meanScore |
|---|---|
0.399999618530273 |
0.521904373168945 |
Mutate
The mutate execution mode extends the stats mode with an important side effect: updating the named graph with a new node property containing the centrality for that node.
The name of the new property is specified using the mandatory configuration parameter mutateProperty.
The result is a single summary row, similar to stats, but with some additional metrics.
The mutate mode is especially useful when multiple algorithms are used in conjunction.
For more details on the mutate mode in general, see Mutate.
mutate mode:CALL gds.closeness.mutate('myGraph', { mutateProperty: 'centrality' })
YIELD centralityDistribution, nodePropertiesWritten
RETURN centralityDistribution.min AS minimumScore, centralityDistribution.mean AS meanScore, nodePropertiesWritten| minimumScore | meanScore | nodePropertiesWritten |
|---|---|---|
0.399999618530273 |
0.521904373168945 |
5 |
Write
The write execution mode extends the stats mode with an important side effect: writing the centrality for each node as a property to the Neo4j database.
The name of the new property is specified using the mandatory configuration parameter writeProperty.
The result is a single summary row, similar to stats, but with some additional metrics.
The write mode enables directly persisting the results to the database.
For more details on the write mode in general, see Write.
write mode:CALL gds.closeness.write('myGraph', { writeProperty: 'centrality' })
YIELD centralityDistribution, nodePropertiesWritten
RETURN centralityDistribution.min AS minimumScore, centralityDistribution.mean AS meanScore, nodePropertiesWritten| minimumScore | meanScore | nodePropertiesWritten |
|---|---|---|
0.399999618530273 |
0.521904373168945 |
5 |